Luxembourg's national laboratory (Laboratoire National de Santé - LNS) confirmed to Chronicle.lu that a total of 28,589 SARS-CoV-2 (COVID-19) specimens were sequenced in 2021, of which 25,667 were from Luxembourg residents.
Genomic sequencing is used to identify COVID-19. It is considered particularly useful in determining different variants of the virus. Depending on changes to the genome (i.e. an organism's complete set of RNA / DNA), new variants or lineages (sub-groups of the same variant) are then characterised. Each variant is considerably different from the other and this has various clinical or epidemiological consequences (e.g. regarding transmissibility, infection rates or severity).
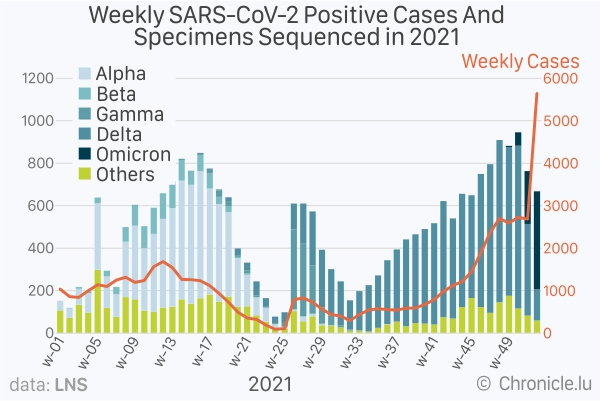
As shown in the graph above, the number of each Variant of Concern (VoC), namely, Alpha, Beta, Gamma, Delta and Omicron, is shown along with “others”, which are sequences either not belonging to any VoC or the data quality is insufficient to be correctly assigned to a VoC. The number of new cases detected each week throughout 2021 is also shown. Note the difference in the y-axis values on the left-hand side (for sequencing) and on the right-hand side (for the number of new cases).
Since it remains important to continuously monitor new emerging variants as well as their weekly changes to allow for adaptations to public health and safety measures, COVID-19 sequencing remains an indispensable tool.
The two other methods more commonly used to determine the presence of COVID-19 in a sample are PCR and rapid antigen tests. The PCR method amplifies only small parts of the virus genome. Therefore, with a PCR test, it is possible to determine if the virus is present in a sample collected from the nose (nasopharyngeal swabs) or throat (oropharyngeal swabs). However, individual variants cannot be ascertained due to the lack of full-length genome sequence. In the case of rapid antigen tests, the antigen (a protein) recognises the spike protein on the outer surface of the SARS-CoV-2 virus. No RNA or DNA sequence is involved, and hence again, individual variants cannot be ascertained. Antigen tests are generally fast (five to fifteen minutes), require minimal training or infrastructure and are much cheaper than PCR or sequencing techniques.
The National Reference Laboratory for Acute Respiratory Infections at LNS in Dudelange is the only site in Luxembourg where all the sequencing related to COVID-19 is analysed. The department receives SARS-CoV-2 positive samples (nasopharyngeal or oropharyngeal swabs analysed by PCR) from the national network of laboratories, including: all specimens from hospital cases; all specimens from post-vaccination cases; specimens from clusters with high transmission; and representative samples of community cases.
The representative samples of community cases are a systematic selection from the weekly COVID-19 positive cases registered in Luxembourg and are selected according to the European Centre for Disease Prevention and Control (ECDC) guidelines.
The positive samples received and processed for sequencing at LNS takes additional time and hence the reporting is usually delayed by two weeks from the date of sample collection.
In 2021, a total of 58,579 COVID-19 cases were detected among Luxembourg residents, of which 25,667 resident samples were sequenced – representative of an estimated 23,900 people, equivalent to 40.8% of all cases.
The difference between the number of resident samples and the number of people is due to the fact that, at times, several specimens were collected from one person, like in the case of reinfections or hospitalisations.
The Omicron VoC, first detected in five specimens (7.9%) from week 49, increased to 462 specimens representing 21.9% of all the circulating lineages in Luxembourg by week 52, the last week of 2021.
As of the latest data from LNS for week 4 of 2022 (24 to 30 January), Omicron VoC represented 99.5% of all circulating lineages in Luxembourg, as determined by the sequencing of 924 resident specimens.








